Usage
Get started
Navigate to the Pipelines page and select the Launch button under the RNA-seq card.
Requestor information
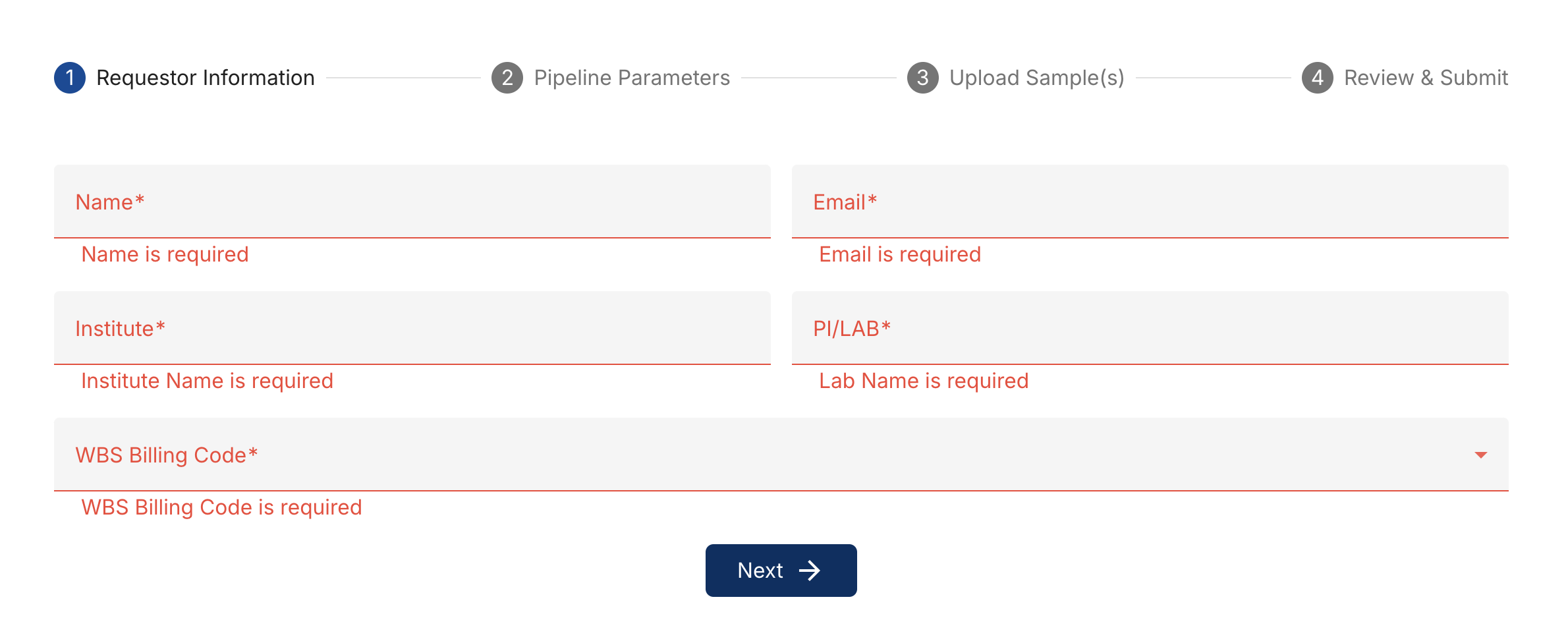
Fill in your full name, email address, institute, principal investigator name, and WBS billing code. This information just lets us know who and which account should be invoiced. Don't worry, you will not be invoiced for anything until the pipeline has completed successfully and you have the results you need.
Parameters
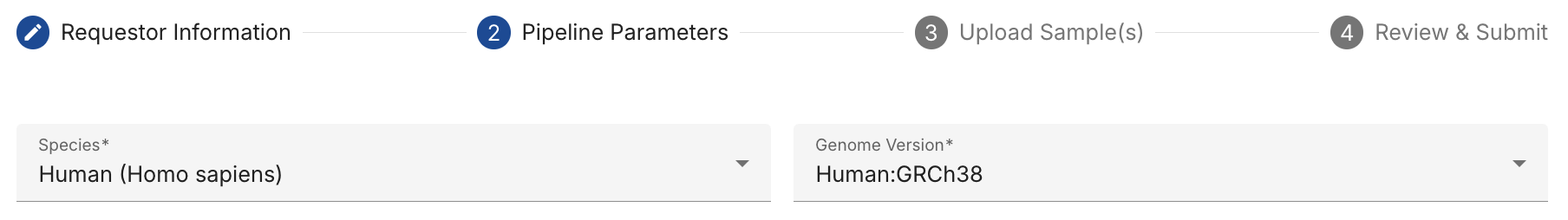
Select the appropriate reference genome and version to use for sequence alignment and transcript quantification.
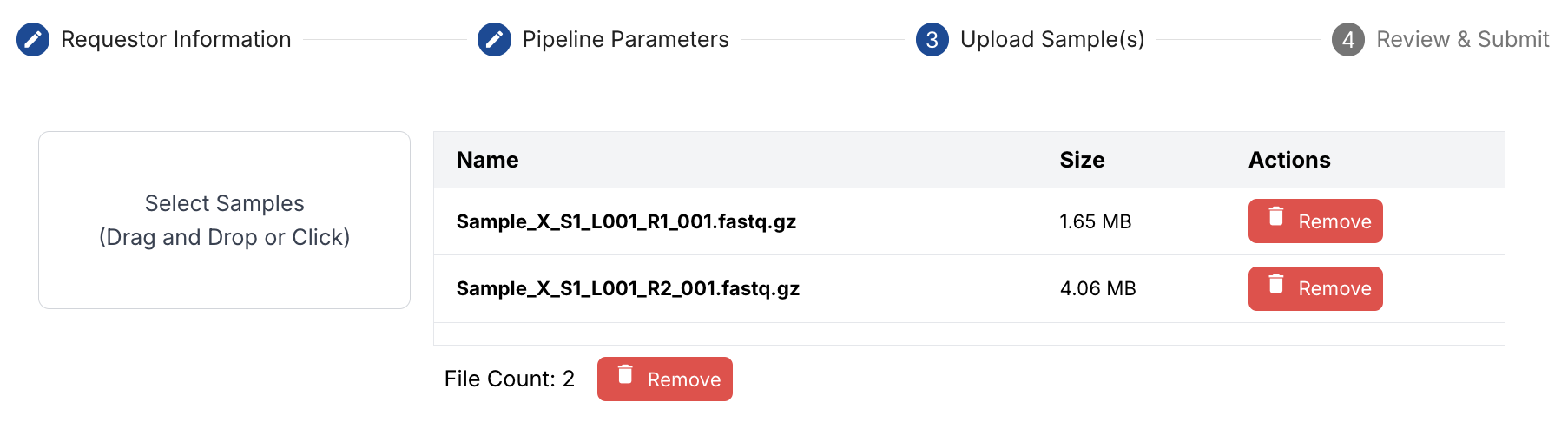
Upload samples
Simply drag-and-drop the FASTQ files from your computer onto the dashboard.
Sample sheet
A sample sheet containing basic information about your experimental design is also required. The header section contains three pieces of information: sample, a unique label for the sample being processed; fastq_1, file name of the forward reads; fastq_2, file name of the reverse reads.
sample,fastq_1,fastq_2
Treatment_A1,Treatment_A1_S1_L001_R1_001.fastq.gz,Treatment_A1_S1_L001_R2_001.fastq.gz
Treatment_A2,Treatment_A2_S2_L001_R1_001.fastq.gz,Treatment_A2_S2_L001_R2_001.fastq.gz
Treatment_A3,Treatment_A3_S3_L001_R1_001.fastq.gz,Treatment_A3_S3_L001_R2_001.fastq.gz
Control_A1,Control_A1_S4_L001_R1_001.fastq.gz,Treatment_A1_S4_L001_R2_001.fastq.gz
Control_A2,Control_A2_S5_L001_R1_001.fastq.gz,Treatment_A2_S5_L001_R2_001.fastq.gz
Control_A3,Control_A3_S6_L001_R1_001.fastq.gz,Treatment_A3_S6_L001_R2_001.fastq.gz
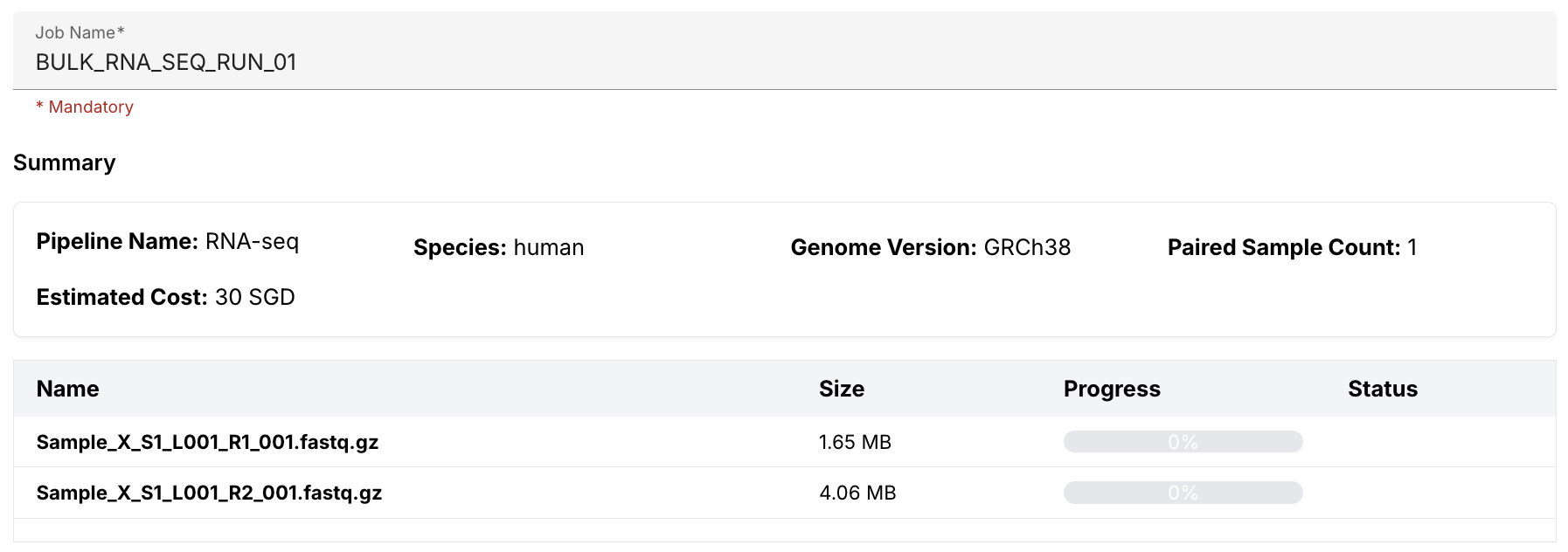
Review and submit
Once the samples have been uploaded, your are ready to submit your job. All you need to do is assign the run a unique job name; review the Summary section to ensure everything looks correct; and then submit your job.